Scale-invariant minimum mean square error (MMSE) improper uniform prior estimators of transformed amplitude
This answer presents a family scale-invariant estimators, parameterized by a single parameter which controls both the Bayesian prior distribution of amplitude and the transformation of amplitude to another scale. The estimators are minimum mean square error (MMSE) estimators in the transformed amplitude scale. An improper uniform prior of transformed amplitude is assumed. Available transformations include a linear scale (no transformation) and can approach a logarithmic scale whereby the estimator approaches zero everywhere. The estimators can be parameterized to attain low sum of square error at negative signal-to-noise ratios (SNRs).
Bayesian estimation
The maximum likelihood (ML) estimator in my first answer performed rather poorly. The ML estimator can also be understood as a Bayesian maximum a posteriori (MAP) estimator given an improper uniform prior probability distribution. Here, improper means that the prior extends from zero to infinity with infinitesimal density. Because the density is not a real number, the prior is not a proper distribution, but it may still give a proper posterior distribution by Bayes' theorem which can then be used to obtain a MAP or an MMSE estimate.
They Bayes' theorem in terms of probability density functions (PDFs) is:
$$\operatorname{PDF}(a\mid m) = \frac{\operatorname{PDF}(m\mid a)\,\operatorname{PDF}(a)}{\operatorname{PDF}(m)} = \frac{\operatorname{PDF}(m\mid a)\,\operatorname{PDF}(a)}{\int_0^\infty\operatorname{PDF}(m\mid a)\,\operatorname{PDF}(a)\,da}.\tag{1}$$
A MAP estimator $\hat a_\text{MAP}$ is the argument of the posterior PDF that maximizes it:
$$\hat a_\text{MAP} = \underset{a}{\operatorname{arg\,max}}\operatorname{PDF}(a \mid m).\tag{2}$$
An MMSE estimator $\hat a_\text{MMSE}$ is the posterior mean:
$$\hat a_\text{MMSE} = \underset{\hat a}{\operatorname{arg\,max}}\operatorname{E}[(a - \hat a)^2\mid m] = \operatorname{E}[a\mid m] = \int_0^\infty a \operatorname{PDF}(a\mid m)da.\tag{3}$$
An improper uniform prior is not the only scale-invariant prior. Any prior PDF satisfying:
$$\operatorname{PDF(|X_k|)} \propto |X_k|^{\varepsilon-1},\tag{4}$$
with real exponent $\varepsilon-1,$ and $\propto$ meaning: "is proportional to", is scale-invariant in the sense that the product of $X_k$ and a positive constant still follows the same distribution (see Lauwers et al. 2010).
A family of estimators
A family of estimators shall be presented, with these properties:
- Scale-invariance: If the complex clean bin $X_k,$ or equivalently the clean amplitude $|X_k|,$ and the noise standard deviation $\sigma$ are each multiplied by the same positive constant, then also the estimated amplitude $\widehat{|X_k|}$ gets multiplied by that constant.
- Minimum mean square transformed-amplitude error.
- Improper uniform prior of transformed amplitude.
We shall use normalized notation:
$$\begin{array}{ll} a &= \frac{|X_k|}{\sigma}&\text{normalized clean amplitude,}\\ m &= \frac{|Y_k|}{\sigma}&\text{normalized noisy magnitude,}\\ 1 &= \left(\frac{\sigma}{\sigma}\right)^2&\text{normalized variance of noise,}\\ \mathrm{SNR} &= \left(\frac{|X_k|}{\sigma}\right)^2 = a^2&\text{signal-to-noise ratio ($10\log_{10}(\mathrm{SNR})$ dB),}\end{array}\tag{5}$$
where $|X_k|$ is the clean amplitude we wish to estimate from the noisy magnitude $|Y_k|$ of bin value $Y_k$ whicy equals the sum of clean bin value $X_k$ plus circularly symmetric complex Gaussian noise of variance $\sigma^2.$ The scale-invariant prior of $|X_k|$ given in Eq. 4 is carried over to the normalized notation as:
$$\operatorname{PDF}(a) \propto a^{\varepsilon - 1}.\tag{6}$$
Let $g(a)$ be an increasing transformation function of amplitude $a.$ The improper uniform prior of transformed amplitude is denoted by:
$$\operatorname{PDF}\big(g(a)\big) \propto 1.\tag{7}$$
Eqs. 6 & 7 together determine the family of possible amplitude transformations. They are related by a change of variables:
$$\begin{array}{rrcl}&g'(a) \operatorname{PDF}\big(g(a)\big) &=& \operatorname{PDF}(a)\\ \displaystyle\Rightarrow&\quad g'(a) &\propto& a^{\varepsilon - 1}\\ \Rightarrow&g(a) &\propto& \displaystyle\int a^{\varepsilon - 1} da = \frac{a^\varepsilon}{\varepsilon} + c\\ \Rightarrow&g(a) &=& \displaystyle\frac{c_1a^\varepsilon}{\varepsilon} + c_0.\end{array}\tag{8}$$
We assume without proof that the choice of the constants $c_0$ and $c_1$ will not affect the amplitude estimate. For convenience we set:
$$\begin{array}{rc}&g(1) = 1\quad\text{and}\quad g'(1) = 1\\ \Rightarrow&c_0 = \displaystyle\frac{\varepsilon - 1}{\varepsilon}\quad\text{and}\quad c_1 = 1\\ \Rightarrow&g(a) = \displaystyle\frac{a^\varepsilon + \varepsilon - 1}{\varepsilon},\\ \end{array}\tag{9}$$
which has a special linear case:
$$g(a) = a\quad\text{if}\quad \varepsilon = 1,\tag{10}$$
and a limit:
$$\lim_{\varepsilon \to 0}g(a) = \log(a) + 1.\tag{11}$$
The transformation function can conveniently represent the linear amplitude scale (at $\varepsilon = 1$) and can approach a logarithmic amplitude scale (as $\varepsilon \to 0$). For positive $\varepsilon,$ the support of the PDF of transformed amplitude is:
$$\begin{eqnarray}&0 < a < \infty&\\ \Rightarrow\quad&\frac{\varepsilon - 1}{\varepsilon} < g(a) < \infty,&\end{eqnarray}\tag{12}$$
The inverse transformation function is:
$$g^{-1}\big(g(a)\big) = \big(\varepsilon g(a) - \varepsilon + 1\big)^{1/\varepsilon} = a.\tag{13}$$
The transformed estimate is then, using the law of the unconscious statistician:
$$\begin{gather}\hat a_\text{uni-MMSE-xform} = \underset{\hat a}{\operatorname{arg\,min}}\operatorname{E}\left[\big(g(a) - g(\hat a)\big)^2\mid m\right] = g^{-1}\big(\operatorname{E}[g(a) \mid m]\big)\\ = g^{-1}\left(\int_0^\infty g(a) \operatorname{PDF}(a \mid m)\,da\right)\\ = g^{-1}\left(\frac{\int_0^\infty g(a) f(a \mid m)da}{\int_0^\infty f(a \mid m)da}\right),\end{gather}\tag{14}$$
where $\operatorname{PDF}(a \mid b)$ is the posterior PDF and $f(a \mid m)$ is an unnormalized posterior PDF defined using Bayes' theorem (Eq. 1), the Rician $\operatorname{PDF}(m \mid a) = 2me^{-\left(m^2 + a^2\right)}I_0(2ma)$ from Eq. 3.2 of my ML estimator answer, and Eq. 6:
$$\begin{eqnarray}\operatorname{PDF}(a\mid m) &\propto& \operatorname{PDF}(m\mid a)\,\operatorname{PDF}(a)\\ &\propto&2me^{-\left(m^2 + a^2\right)}I_0(2ma)\times a^{\varepsilon - 1}\\ &\propto&e^{-a^2}I_0(2ma)\,a^{\varepsilon - 1} = f(a \mid m),\end{eqnarray}\tag{15}$$
from which $\operatorname{PDF}(m)$ was dropped from the Bayes' formula because it is constant over $a.$ Combining Eqs. 14, 9 & 15, solving the integrals in Mathematica, and simplifying, gives:
$$\begin{gather}\hat a_\text{uni-MMSE-xform}=g^{-1}\left(\frac{\int_0^\infty \frac{a^\varepsilon + \varepsilon - 1}{\varepsilon} \times e^{-a^2}I_0(2ma)\,a^{\varepsilon - 1}\,da}{\int_0^\infty e^{-a^2}I_0(2ma)\,a^{\varepsilon - 1}\,da}\right)\\ = \left(\varepsilon\frac{\frac{1}{2\varepsilon}\left(\Gamma(\varepsilon) L_{-\varepsilon}(m^2) + (\varepsilon-1) \Gamma(\varepsilon/2) L_{-\varepsilon/2}(m^2)\right)}{\frac{1}{2} \Gamma(\varepsilon/2) L_{-\varepsilon/2}(m^2)} - \varepsilon + 1\right)^{1/\varepsilon}\\ = \left(\frac{\Gamma(\varepsilon) L_{-\varepsilon}(m^2) + (\varepsilon-1) \Gamma(\varepsilon/2) L_{-\varepsilon/2}(m^2)}{\Gamma(\varepsilon/2) L_{-\varepsilon/2}(m^2)} - \varepsilon + 1\right)^{1/\varepsilon}\\ = \left(\frac{\Gamma(\varepsilon) L_{-\varepsilon}(m^2)}{\Gamma(\varepsilon/2) L_{-\varepsilon/2}(m^2)}\right)^{1/\varepsilon},\end{gather}\tag{16}$$
where $\Gamma$ is the gamma function and $L$ is the Laguerre function. The estimator collapses to zero everywhere as $\varepsilon \to 0,$ so it does not make sense to use negative $\varepsilon,$ which would emphasis small values of $a$ even further and give an improper posterior distribution. Some special cases are:
$$\hat a_\text{uni-MMSE-xform} = \sqrt{m^2 + 1},\quad\text{if }\varepsilon = 2,\tag{17}$$
$$\hat a_\text{uni-MMSE} = \hat a_\text{uni-MMSE-xform}= \frac{e^{m^2/2}}{\sqrt{\pi} I_0(m^2/2)},\quad\text{if }\varepsilon = 1,\tag{18}$$
approximated at large $m$ by (see calculation) a truncated Laurent series:
$$\hat a_\text{uni-MMSE} \approx m - \frac{1}{4m} - \frac{7}{32m^3} - \frac{59}{128m^5},\tag{19}$$
This asymptotic approximation has an absolute maximum amplitude error of less than $10^{-6}$ for $m > 7.7.$
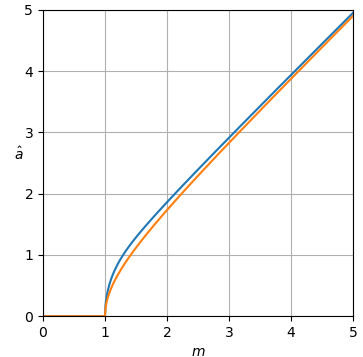
The estimator curves are shown in Fig. 1.

Figure 1. Estimator $\hat a_\text{uni-MMSE-xform}$ as function of $m$ for different values of $\varepsilon,$ from top to bottom: blue: $\varepsilon = 2,$ which minimizes the mean square power error assuming an improper uniform prior of power, orange: $\varepsilon = 1,$ which minimizes the mean square amplitude error assuming an improper uniform prior of amplitude, green: $\varepsilon = \frac{1}{2},$ red: $\varepsilon = \frac{1}{4},$ and purple: $\varepsilon = \frac{1}{8}.$
At $m=0$ the curves are horizontal with value:
$$\hat a_\text{uni-MMSE-xform} = \frac{2^{1 - 1/\varepsilon} \bigg(\Gamma\Big(\frac{1 + \varepsilon}{2}\Big)\bigg)^{1/\varepsilon}}{\pi^{1/(2\varepsilon)}},\quad\text{if }m = 0.\tag{20}$$
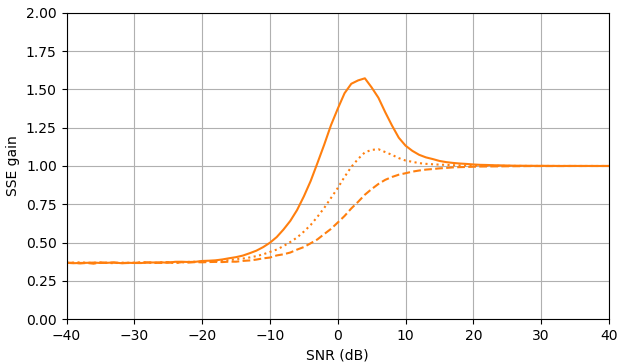
At negative SNR, the uni-MMSE-xform estimator can be parameterized using low $\varepsilon$ to give a lower sum of square error compared to the clamped spectral power subtraction estimator, with a corresponding penalty at intermediate SNR values near 7 dB (Fig. 2).


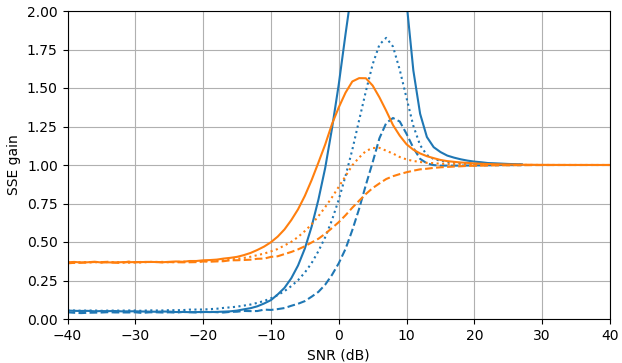
Figure 2. Monte Carlo estimations with a sample size of $10^5,$ of: Solid: gain of sum of square error in estimating $|X_k|$ by $\widehat{|X_k|}$ as compared to estimating it with $|Y_k|,$ dashed: gain of sum of square error in estimating $|X_k|^2$ by $\widehat{|X_k|^2}$ as compared to estimating it with $|Y_k|^2,$ dotted: gain of sum of square error in estimating $X_k$ by $\widehat{|X_k|}e^{i\arg(Y_k)}$ as compared to estimating it with $Y_k.$ Blue: uni-MMSE-xform estimator with $\varepsilon = 1$ (top), $\varepsilon = \frac{1}{2}$ (middle), and $\varepsilon = \frac{1}{4},$ orange: clamped spectral power subtraction.
Python script for Fig. 1
This script extends the question's script A.
def est_a_uni_MMSE_xform(m, epsilon): m = mp.mpf(m) epsilon = mp.mpf(epsilon) if epsilon == 0: return mpf(0) elif epsilon == 1: return mp.exp(m**2/2)/(mp.sqrt(mp.pi)*mp.besseli(0, m**2/2)) elif epsilon == 2: return mp.sqrt(m**2 + 1) else: return (mp.gamma(epsilon)*mp.laguerre(-epsilon, 0, m**2) / (mp.gamma(epsilon/2)*mp.laguerre(-epsilon/2, 0, m**2)))**(1/epsilon) ms = np.arange(0, 6.0625, 0.0625) est_as_uni_MMSE_xform = [[est_a_uni_MMSE_xform(m, 2) for m in ms], [est_a_uni_MMSE_xform(m, 1) for m in ms], [est_a_uni_MMSE_xform(m, 0.5) for m in ms], [est_a_uni_MMSE_xform(m, 0.25) for m in ms], [est_a_uni_MMSE_xform(m, 0.125) for m in ms]] plot_est(ms, est_as_uni_MMSE_xform)
Python script for Fig. 2
This script extends the question's script B. The function est_a_uni_MMSE_xform_fast may be numerically unstable.
from scipy import special def est_a_uni_MMSE_fast(m): return 1/(np.sqrt(np.pi)*special.i0e(m**2/2)) def est_a_uni_MMSE_xform_fast(m, epsilon): if epsilon == 0: return 0 elif epsilon == 1: return 1/(np.sqrt(np.pi)*special.i0e(m**2/2)) elif epsilon == 2: return np.sqrt(m**2 + 1) else: return (special.gamma(epsilon)*special.eval_laguerre(-epsilon, m**2)/(special.gamma(epsilon/2)*special.eval_laguerre(-epsilon/2, m**2)))**(1/epsilon) gains_SSE_a_uni_MMSE = [est_gain_SSE_a(est_a_uni_MMSE_fast, a, 10**5) for a in as_] gains_SSE_a2_uni_MMSE = [est_gain_SSE_a2(est_a_uni_MMSE_fast, a, 10**5) for a in as_] gains_SSE_complex_uni_MMSE = [est_gain_SSE_complex(est_a_uni_MMSE_fast, a, 10**5) for a in as_] plot_gains_SSE(as_dB, [gains_SSE_a_uni_MMSE, gains_SSE_a_sub], [gains_SSE_a2_uni_MMSE, gains_SSE_a2_sub], [gains_SSE_complex_uni_MMSE, gains_SSE_complex_sub]) gains_SSE_a_uni_MMSE_xform_0e5 = [est_gain_SSE_a(lambda m: est_a_uni_MMSE_xform_fast(m, 0.5), a, 10**5) for a in as_] gains_SSE_a2_uni_MMSE_xform_0e5 = [est_gain_SSE_a2(lambda m: est_a_uni_MMSE_xform_fast(m, 0.5), a, 10**5) for a in as_] gains_SSE_complex_uni_MMSE_xform_0e5 = [est_gain_SSE_complex(lambda m: est_a_uni_MMSE_xform_fast(m, 0.5), a, 10**5) for a in as_] plot_gains_SSE(as_dB, [gains_SSE_a_uni_MMSE_xform_0e5, gains_SSE_a_sub], [gains_SSE_a2_uni_MMSE_xform_0e5, gains_SSE_a2_sub], [gains_SSE_complex_uni_MMSE_xform_0e5, gains_SSE_complex_sub]) gains_SSE_a_uni_MMSE_xform_0e25 = [est_gain_SSE_a(lambda m: est_a_uni_MMSE_xform_fast(m, 0.25), a, 10**5) for a in as_] gains_SSE_a2_uni_MMSE_xform_0e25 = [est_gain_SSE_a2(lambda m: est_a_uni_MMSE_xform_fast(m, 0.25), a, 10**5) for a in as_] gains_SSE_complex_uni_MMSE_xform_0e25 = [est_gain_SSE_complex(lambda m: est_a_uni_MMSE_xform_fast(m, 0.25), a, 10**5) for a in as_] plot_gains_SSE(as_dB, [gains_SSE_a_uni_MMSE_xform_0e25, gains_SSE_a_sub], [gains_SSE_a2_uni_MMSE_xform_0e25, gains_SSE_a2_sub], [gains_SSE_complex_uni_MMSE_xform_0e25, gains_SSE_complex_sub])
References
Lieve Lauwers, Kurt Barbe, Wendy Van Moer and Rik Pintelon, Analyzing Rice distributed functional magnetic resonance imaging data: A Bayesian approach, Meas. Sci. Technol. 21 (2010) 115804 (12pp) DOI: 10.1088/0957-0233/21/11/115804.