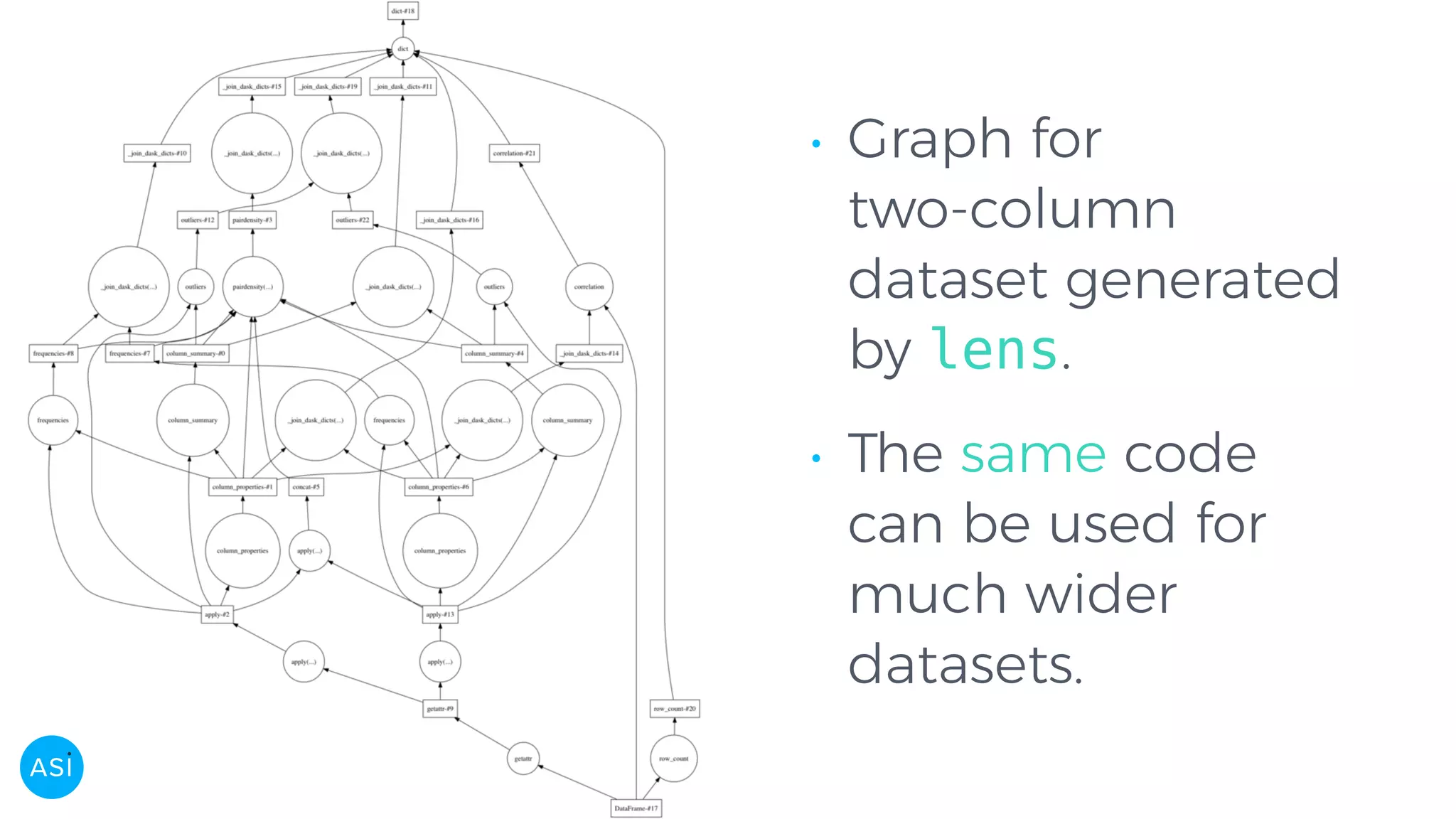
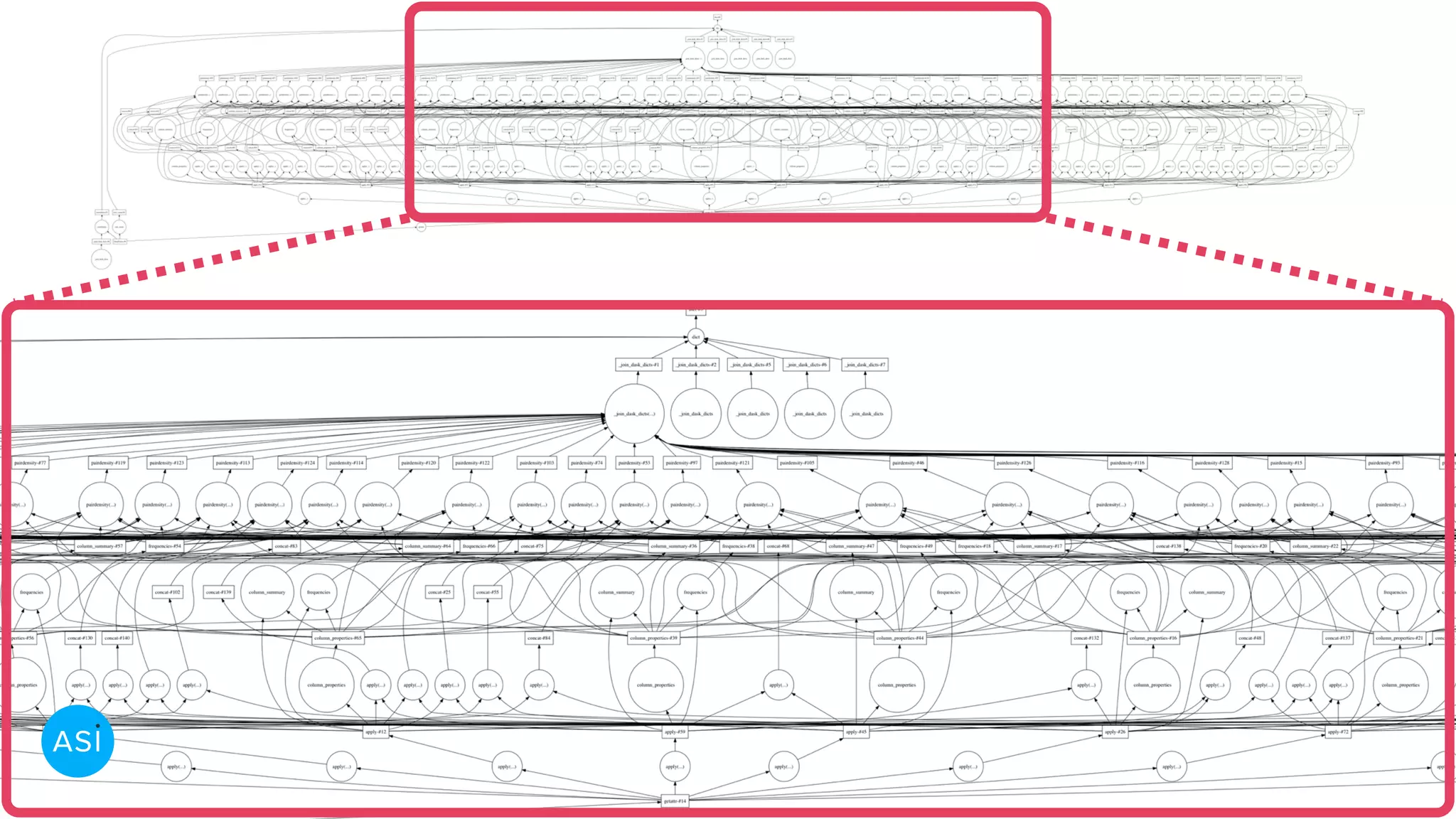
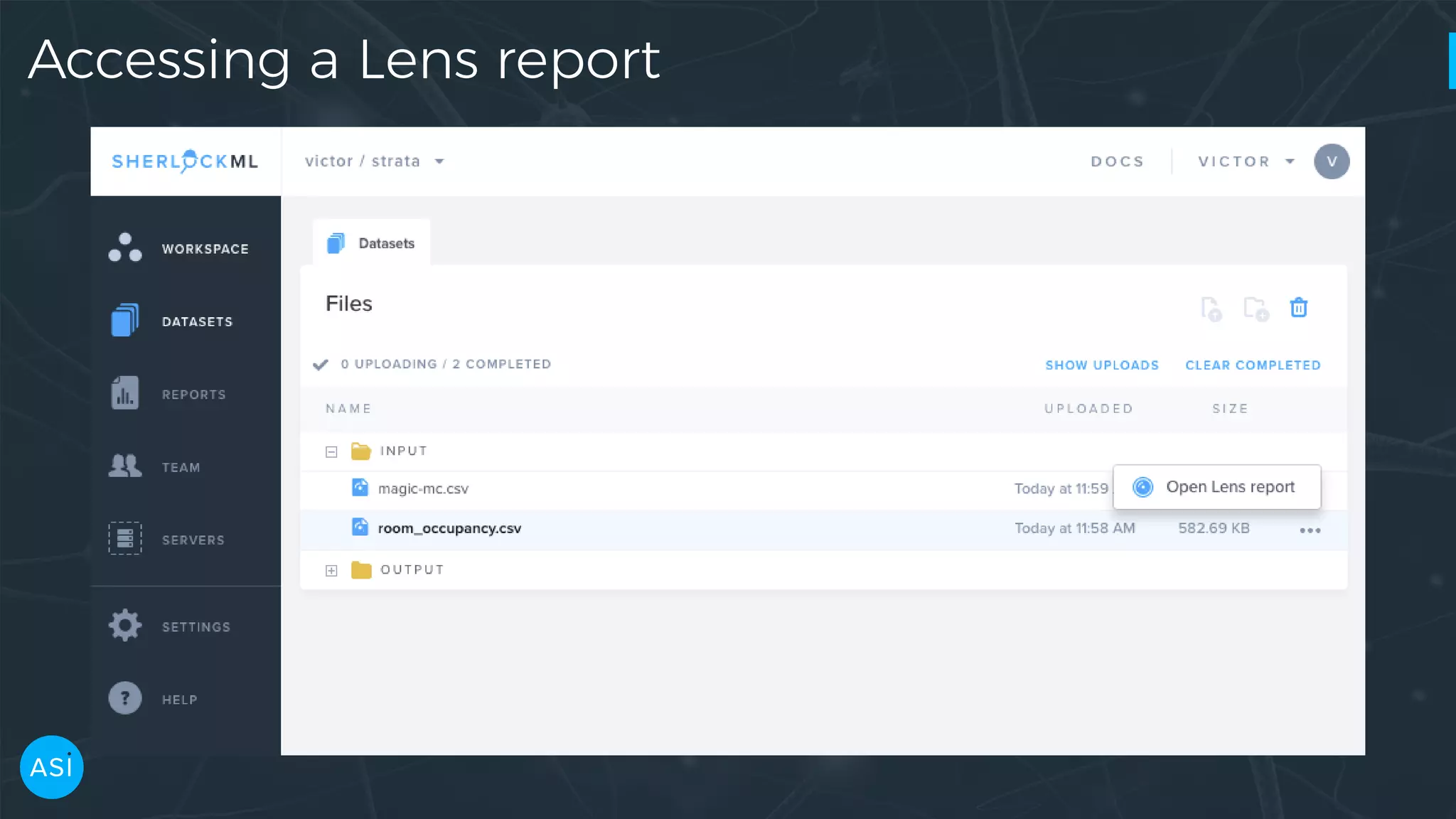
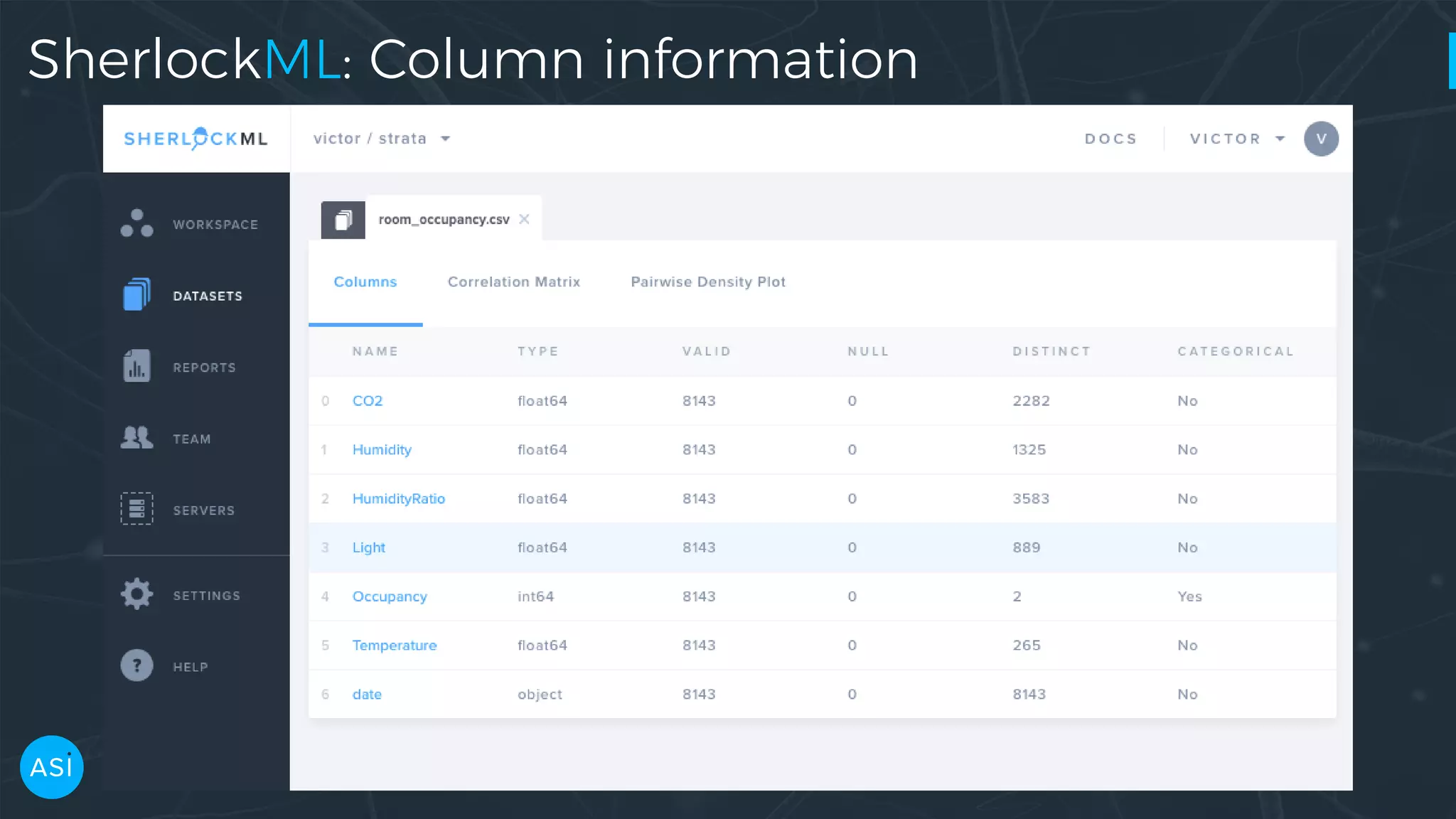
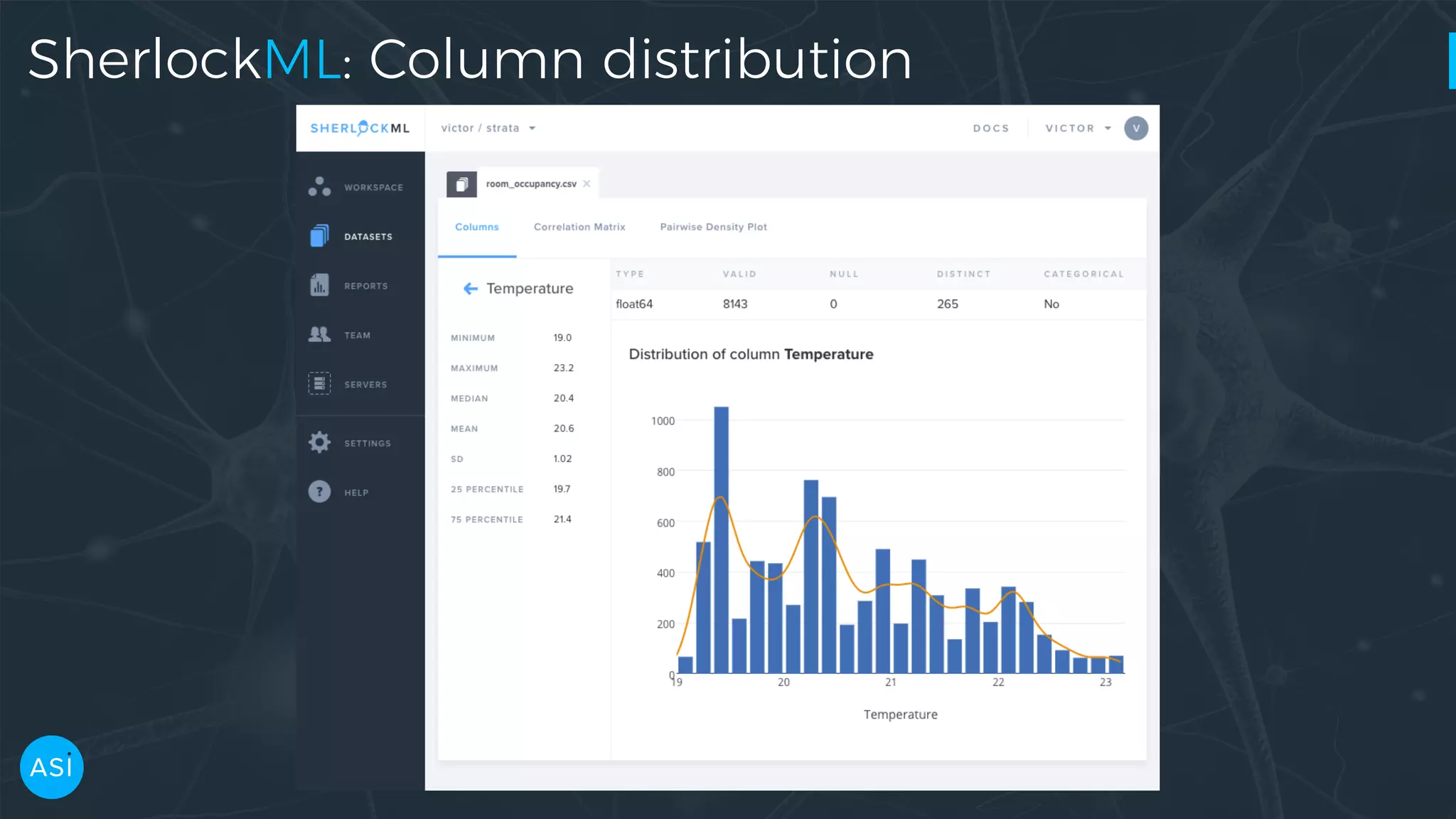
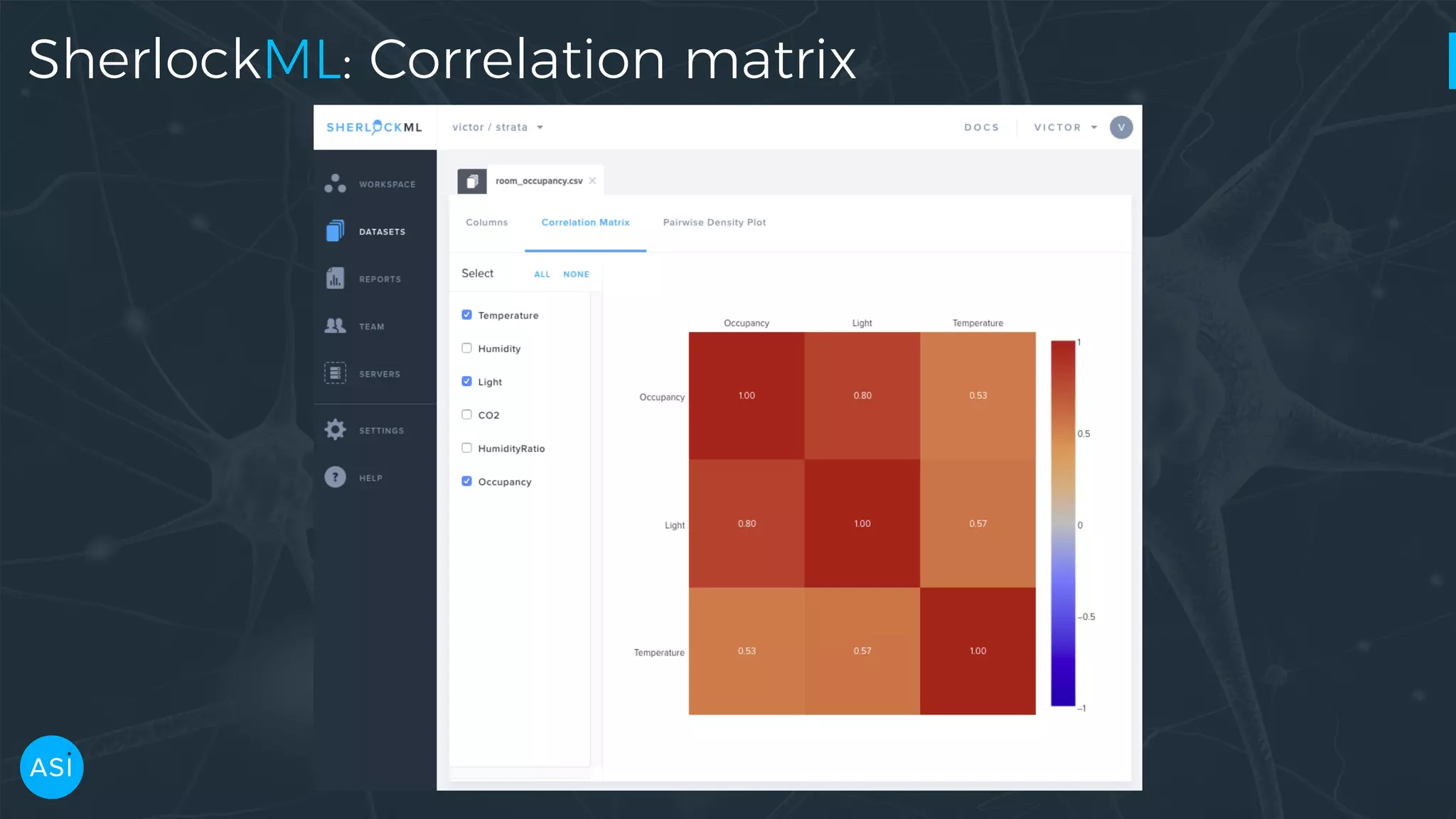
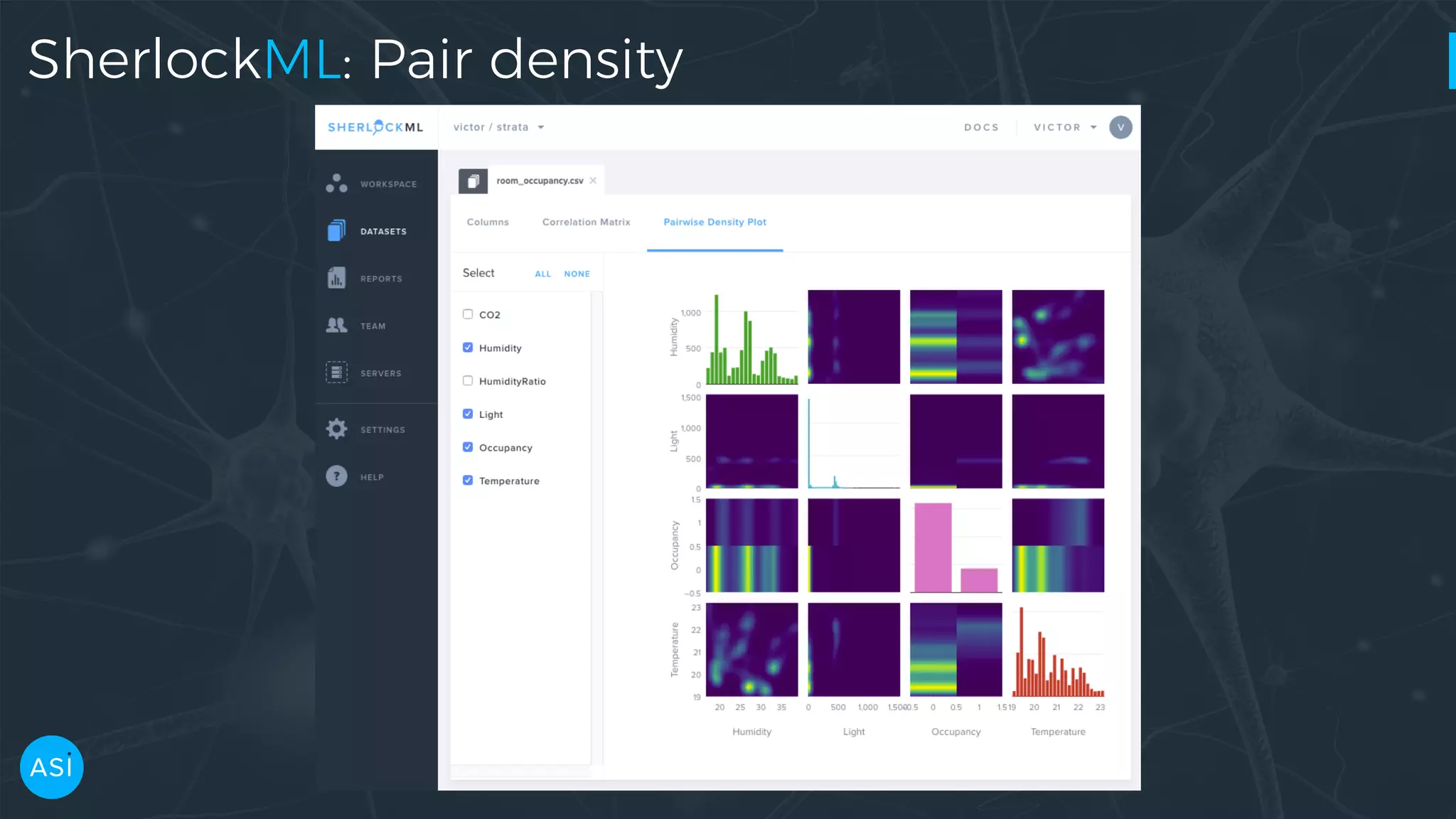
Dask can be used to automate the initial data exploration process by building an analysis graph upfront. Lens is a library and service that uses Dask to compute metrics on datasets like column statistics, correlations, and pair densities. This pre-computes the information needed for exploration so users can interactively examine datasets of any size through Jupyter widgets or a web UI without waiting for new computations. Lens was developed to improve data scientist productivity and is integrated into the SherlockML platform for collaborative data science projects.












![Python interface — Columns >>> ls = lens.Summary.from_json('room_occupancy_lens.json') >>> ls.columns ['date', 'Temperature', 'Humidity', 'Light', 'CO2', 'HumidityRatio', 'Occupancy']](https://image.slidesharecdn.com/victorz-handout-170601083922/75/Automated-Data-Exploration-Building-efficient-analysis-pipelines-with-Dask-13-2048.jpg)
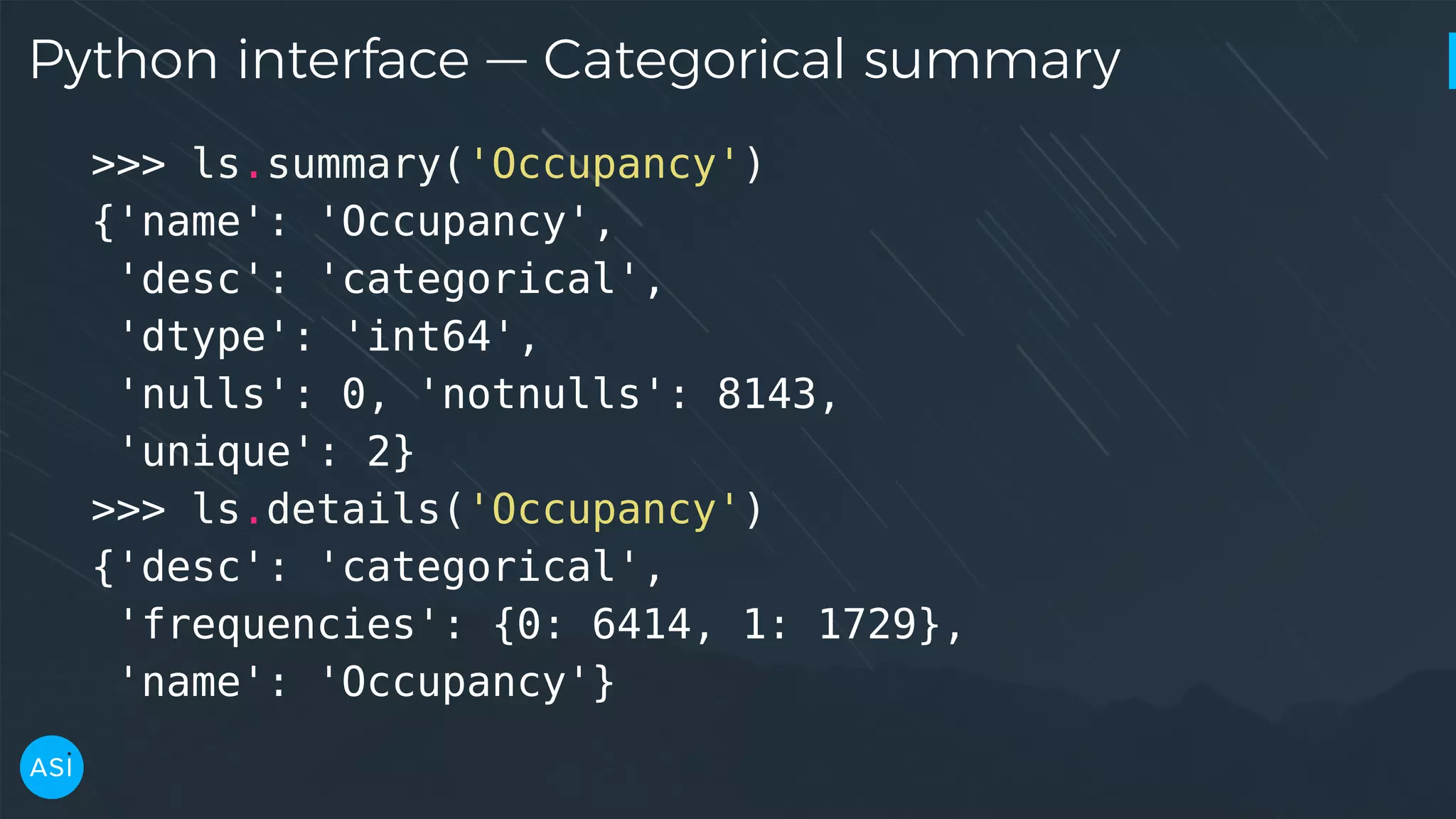
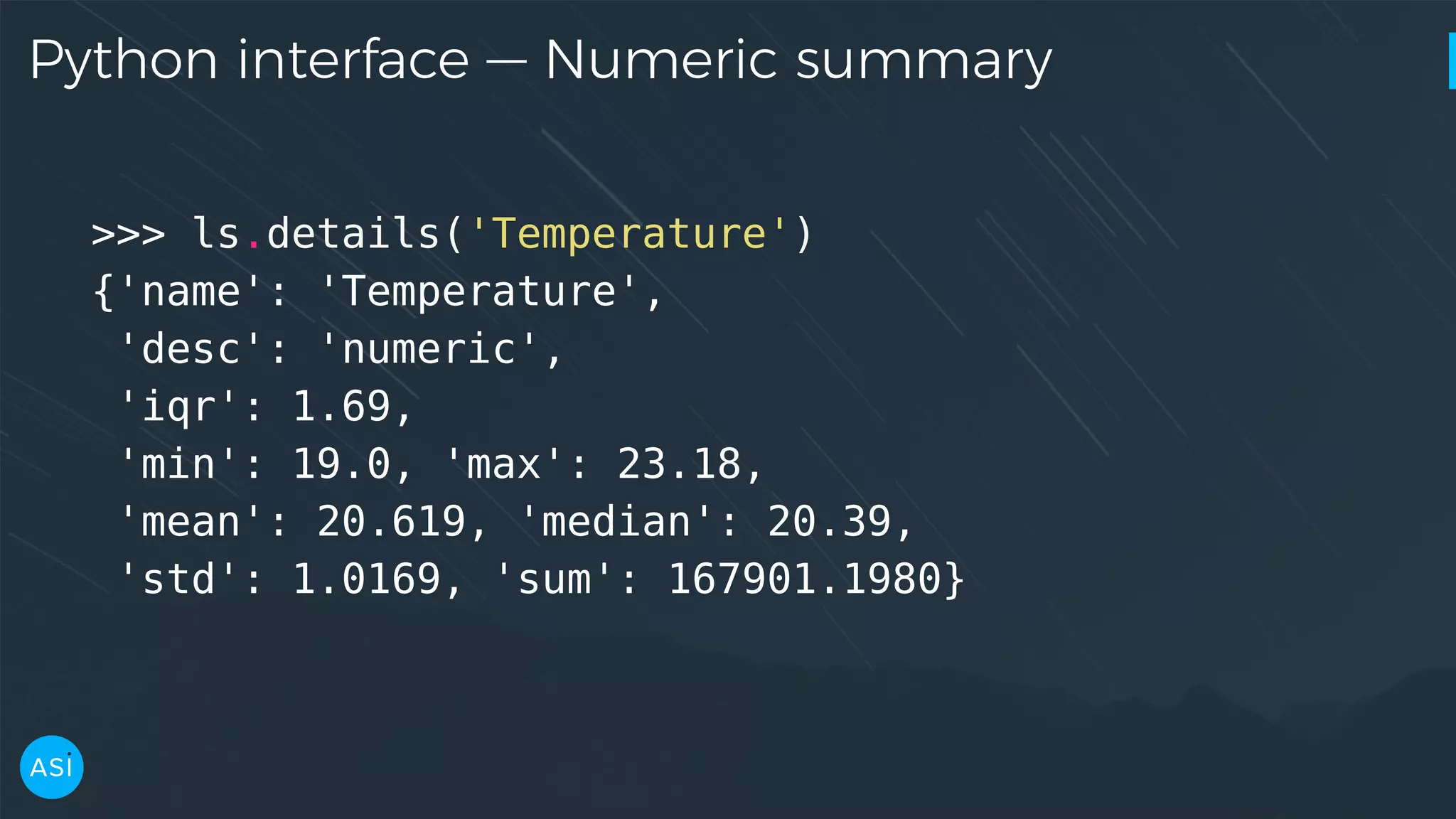
![Python interface — KDE, PDF >>> x, y = ls.kde('Temperature') >>> x[np.argmax(y)] 19.417999999999999 >>> temperature_pdf = ls.pdf('Temperature') >>> temperature_pdf([19, 20, 21, 22]) array([ 0.01754398, 0.76491742, 0.58947765, 0.28421244])](https://image.slidesharecdn.com/victorz-handout-170601083922/75/Automated-Data-Exploration-Building-efficient-analysis-pipelines-with-Dask-16-2048.jpg)


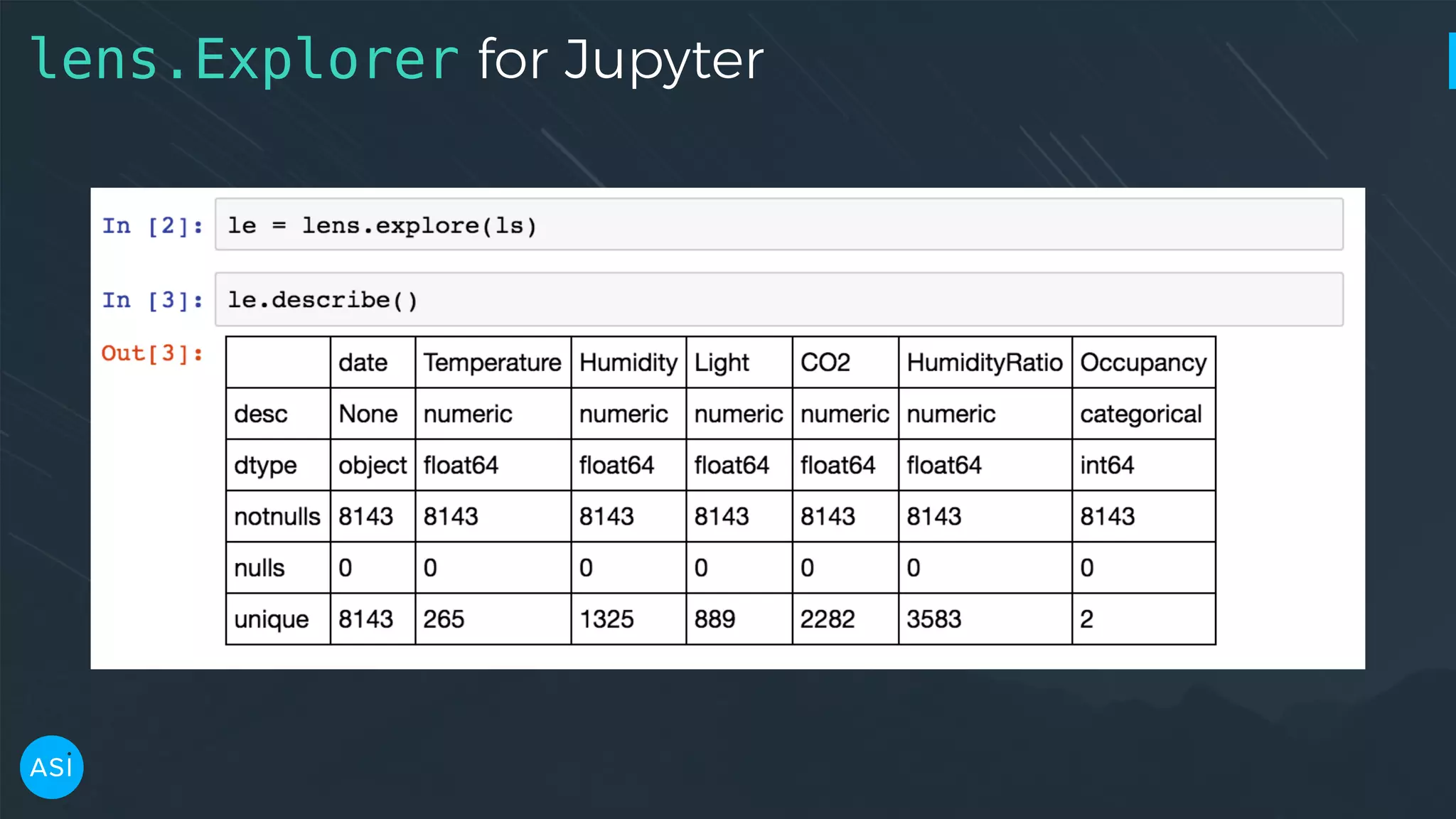
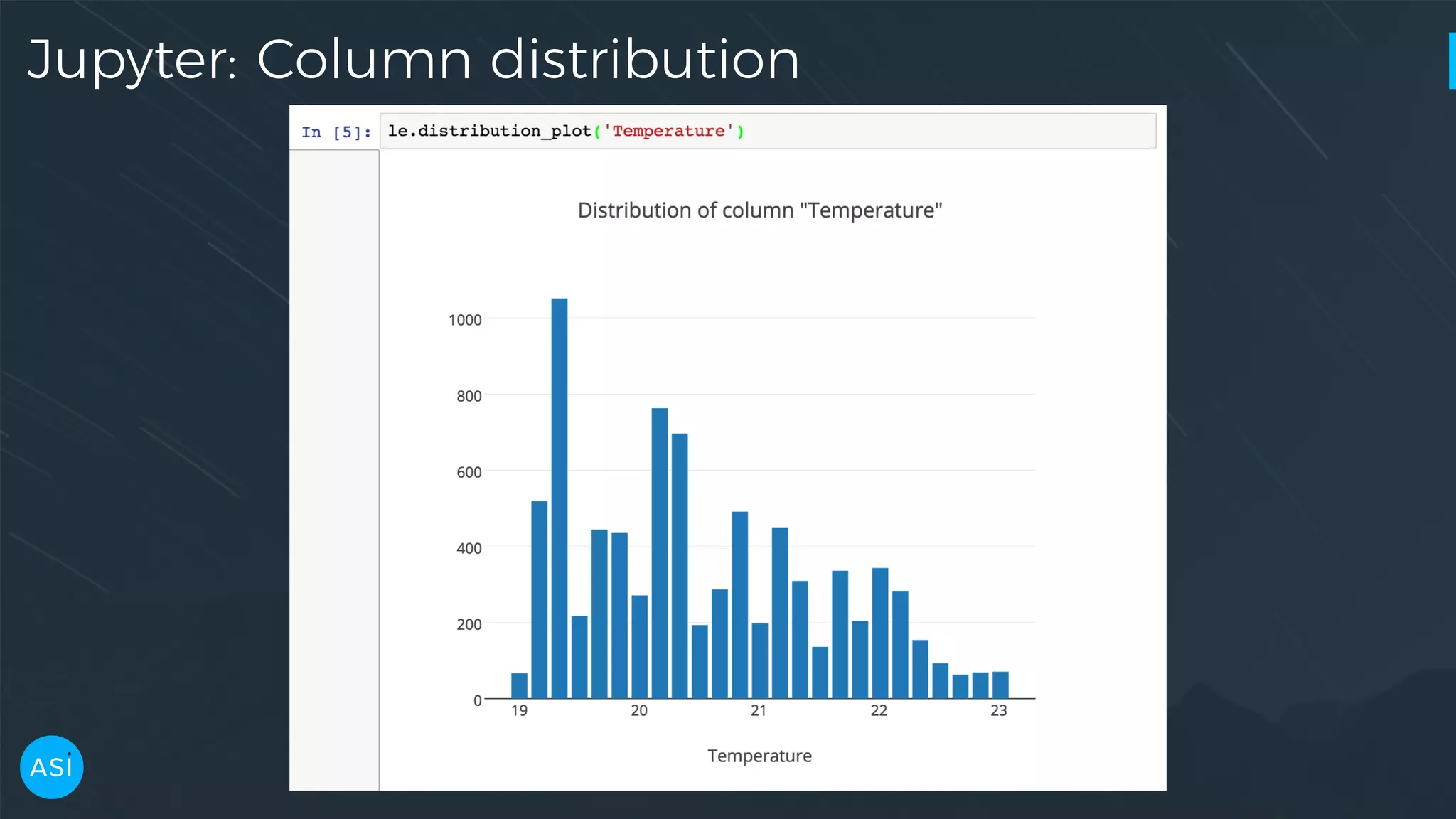
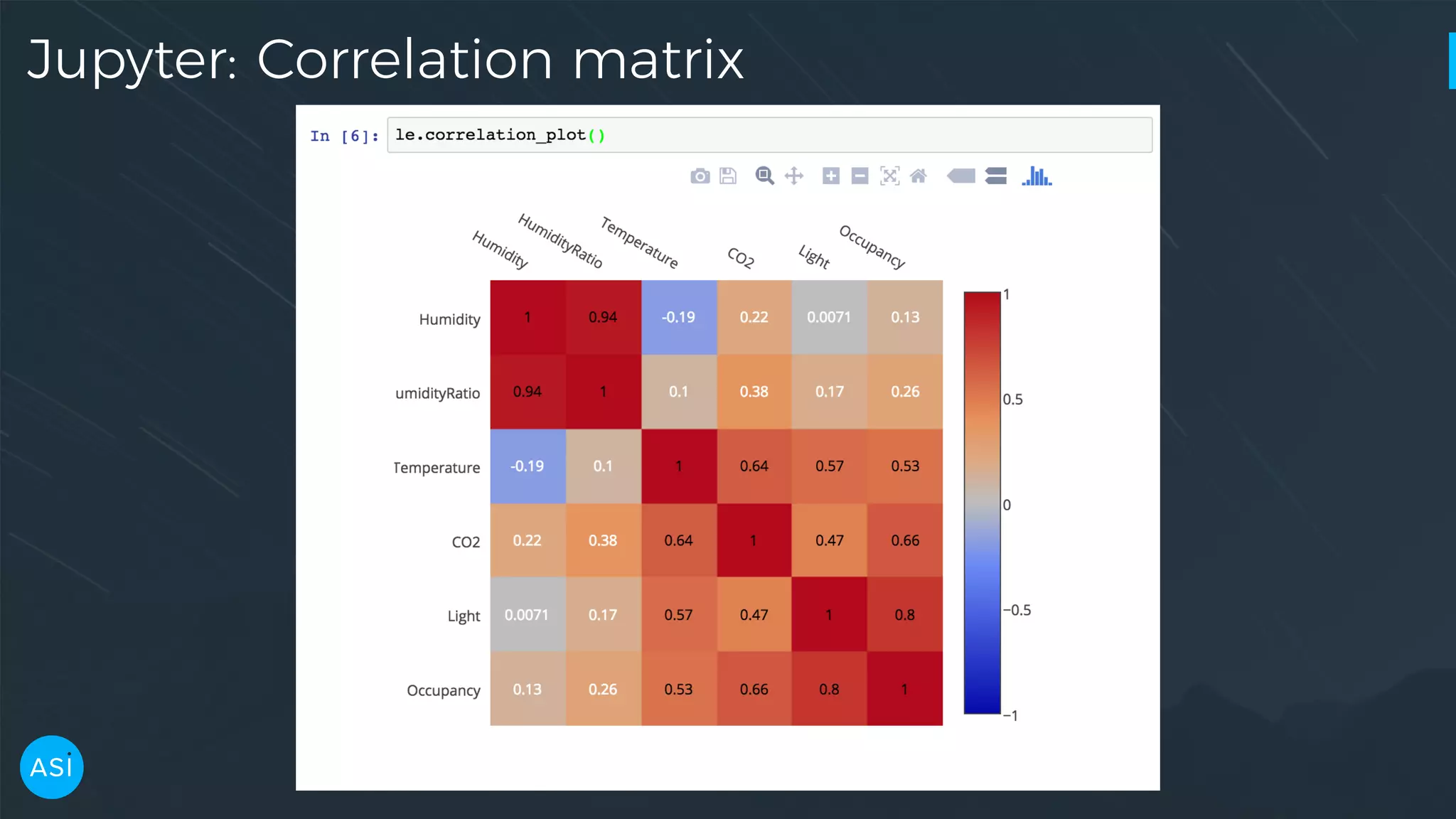









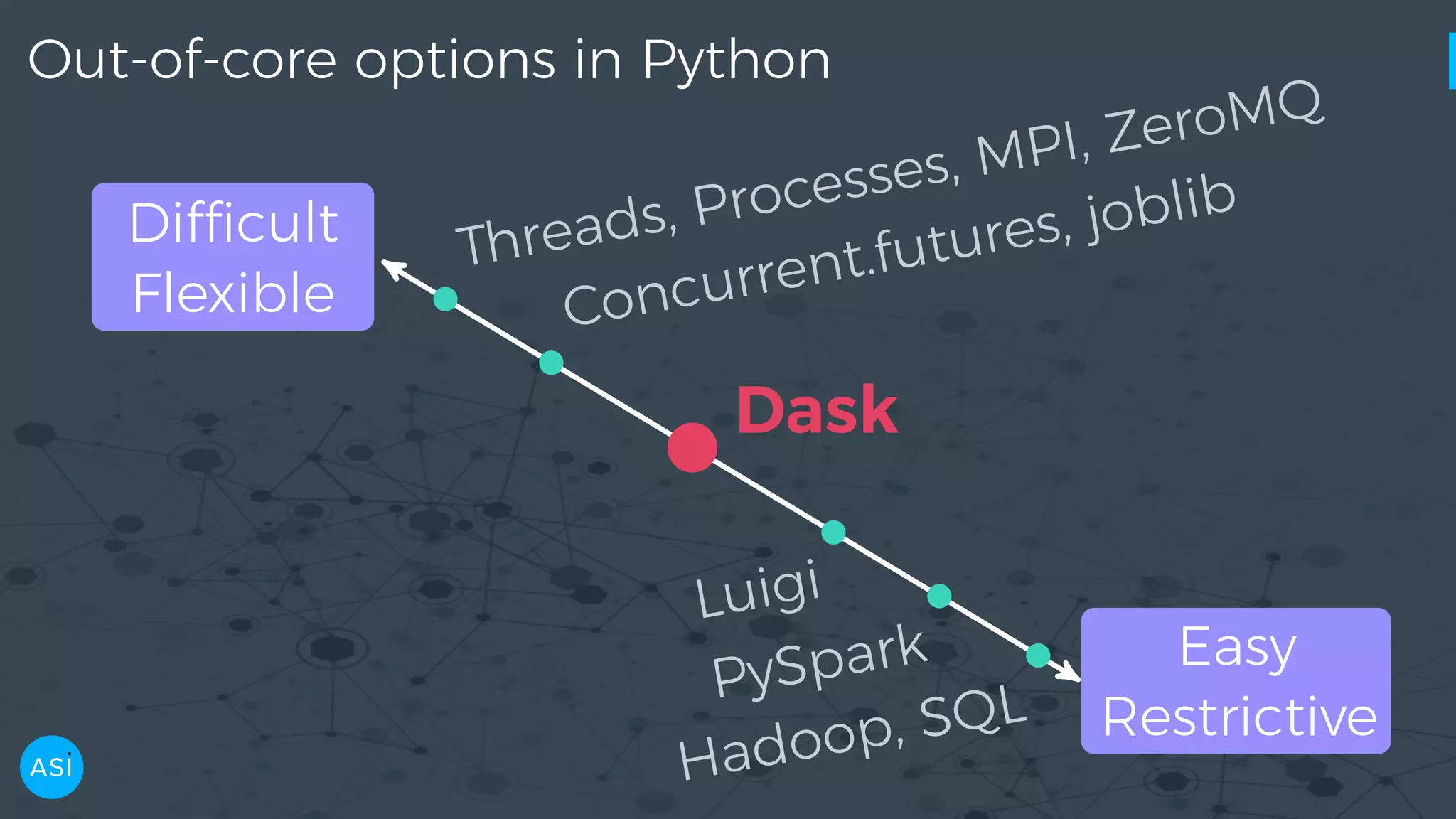



![dask.delayed — Build you own DAG files = ['myfile.a.data', 'myfile.b.data', 'myfile.c.data'] loaded = [load(i) for i in files] cleaned = [clean(i) for i in loaded] analyzed = analyze(cleaned) store(analyzed)](https://image.slidesharecdn.com/victorz-handout-170601083922/75/Automated-Data-Exploration-Building-efficient-analysis-pipelines-with-Dask-39-2048.jpg)

![dask.delayed — Build you own DAG files = ['myfile.a.data', 'myfile.b.data', 'myfile.c.data'] loaded = [load(i) for i in files] cleaned = [clean(i) for i in loaded] analyzed = analyze(cleaned) stored = store(analyzed) clean-2 analyze cleanload-2 analyze store clean-3 clean-1 load storecleanload-1 cleanload-3load load stored.compute()](https://image.slidesharecdn.com/victorz-handout-170601083922/75/Automated-Data-Exploration-Building-efficient-analysis-pipelines-with-Dask-41-2048.jpg)



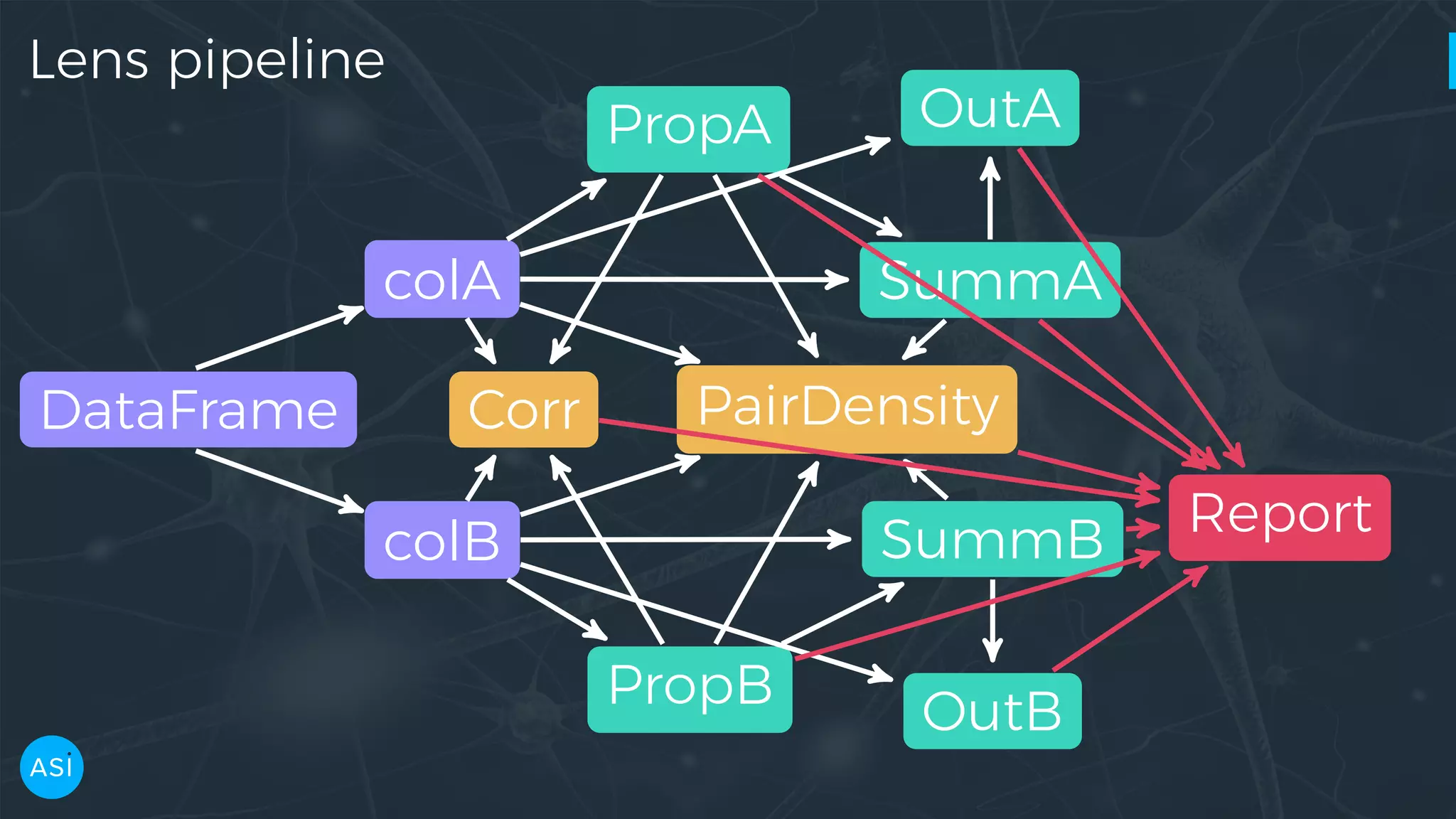
![Building the graph with dask.delayed # Create a series for each column in the DataFrame. columns = df.columns df = delayed(df) cols = {k: delayed(df.get)(k) for k in columns} # Create the delayed reports using Dask. cprops = {k: delayed(metrics.column_properties)(cols[k]) for k in columns} csumms = {k: delayed(metrics.column_summary)(cols[k], cprops[k]) for k in columns} corr = delayed(metrics.correlation)(df, cprops)](https://image.slidesharecdn.com/victorz-handout-170601083922/75/Automated-Data-Exploration-Building-efficient-analysis-pipelines-with-Dask-46-2048.jpg)
![Building the graph with dask.delayed pdens_results = [] if pairdensities: for col1, col2 in itertools.combinations(columns, 2): pdens_df = delayed(pd.concat)([cols[col1], cols[col2]]) pdens_cp = {k: cprops[k] for k in [col1, col2]} pdens_cs = {k: csumms[k] for k in [col1, col2]} pdens_fr = {k: freqs[k] for k in [col1, col2]} pdens = delayed(metrics.pairdensity)( pdens_df, pdens_cp, pdens_cs, pdens_fr) pdens_results.append(pdens) # Join the delayed per-metric reports into a dictionary. report = delayed(dict)(column_properties=cprops, column_summary=csumms, pair_density=pdens_results, ...) return report](https://image.slidesharecdn.com/victorz-handout-170601083922/75/Automated-Data-Exploration-Building-efficient-analysis-pipelines-with-Dask-47-2048.jpg)