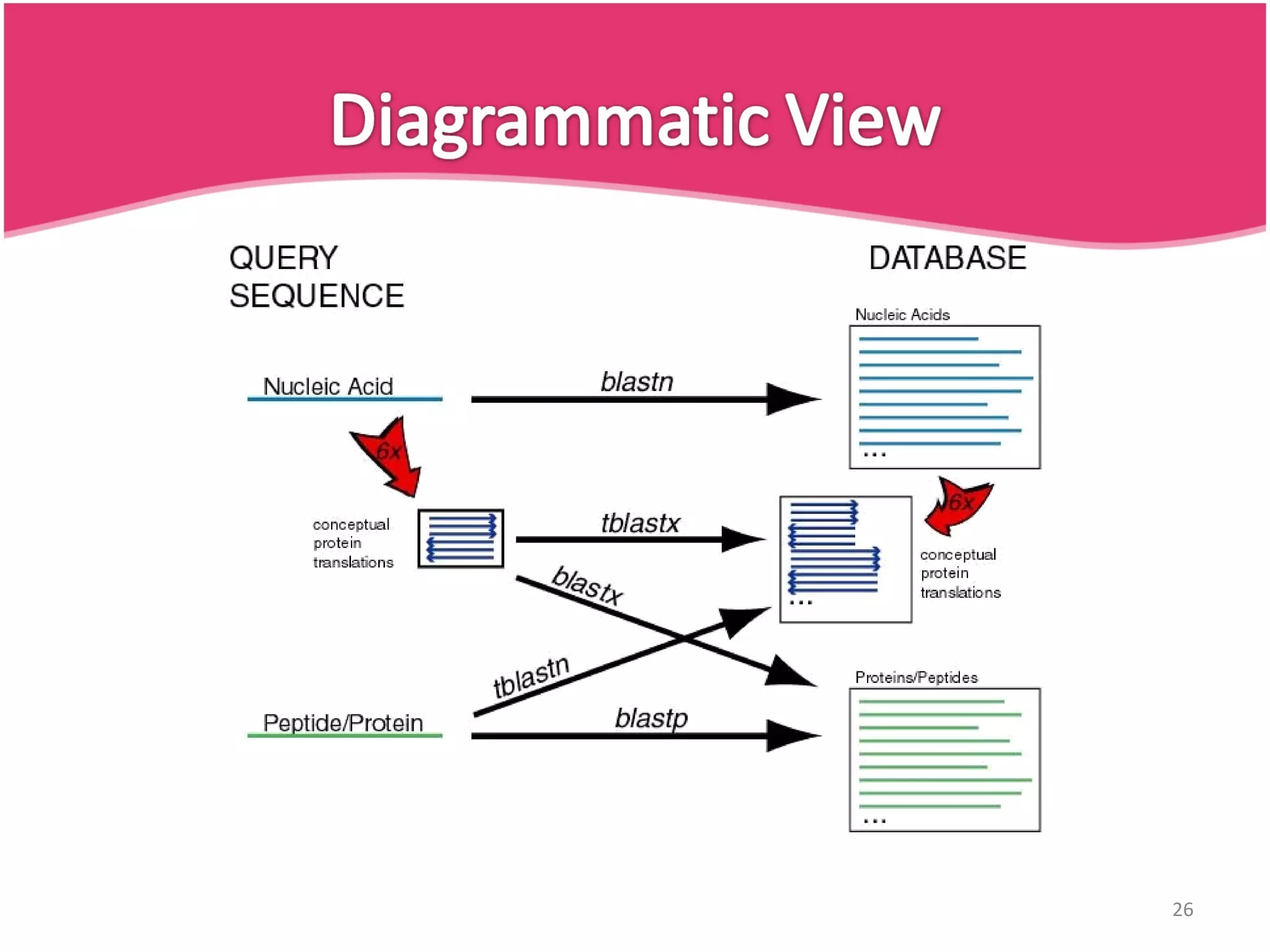
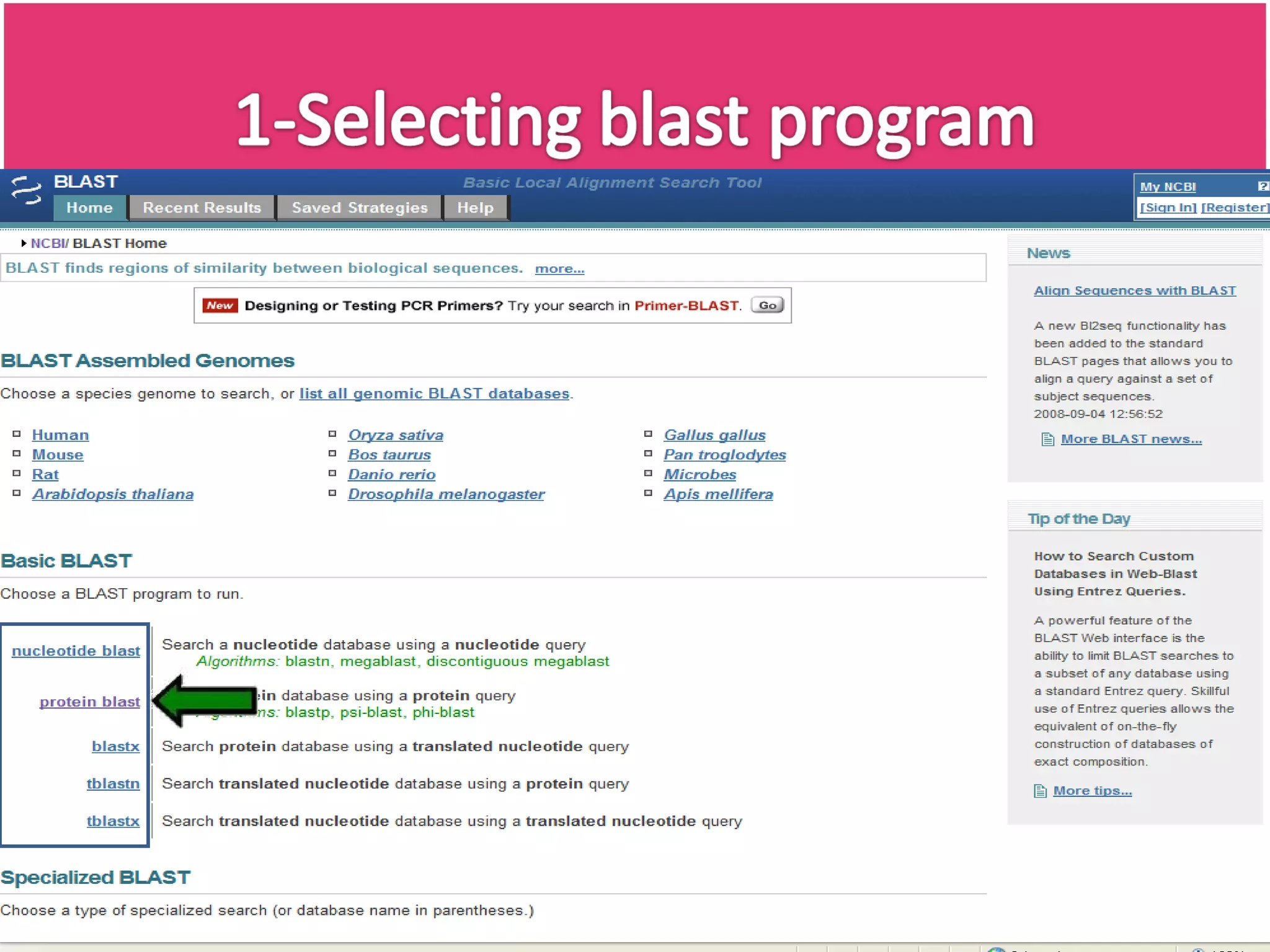
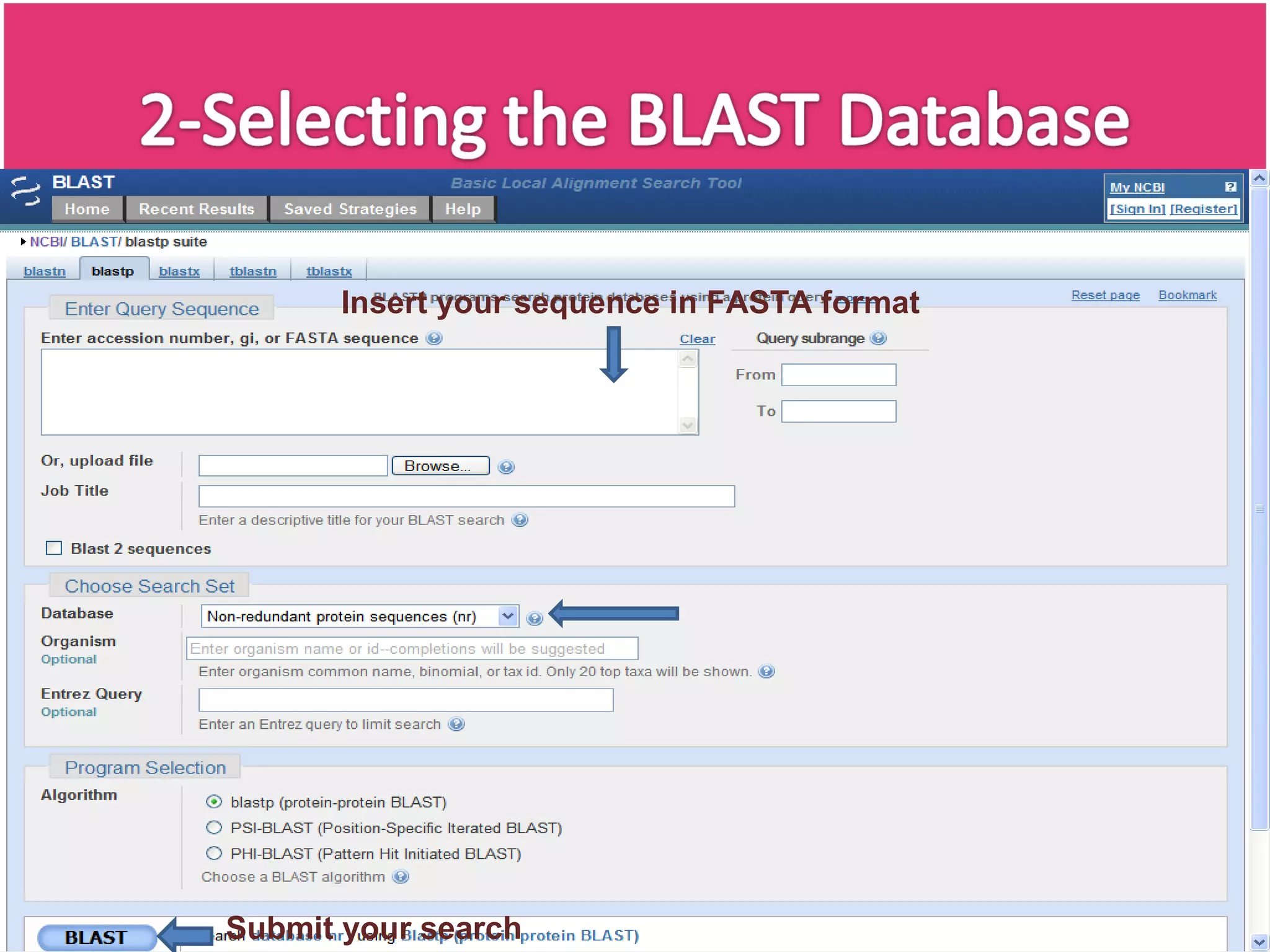
The document discusses the Basic Local Alignment Search Tool (BLAST), a set of algorithms for comparing gene and protein sequences against public databases. It highlights BLAST's efficiency over traditional alignment methods, particularly its speed, and provides an overview of its functionality and various types of BLAST programs. The exponential growth of sequence data and the importance of rapid similarity searching for better understanding of sequences are emphasized.