This repo is designed for a seamless experience of pRF mapping and is closing the gap between preprocessing (using e.g. fmriprep) and the pRF analysis (using github.com/vistalab/PRFmodel).
You can pull the latest docker: docker pull davidlinhardt/prfprepare:1.4.0 Or as singularity image: singularity build prfprepare_1.4.0.sif docker://davidlinhardt/prfprepare:1.4.0
The docker is built up in three stages:
-
build the stimuli from stimulus image and vistadisp log files (if vistadisp was used for stimulus presentation)
-
create a mask in fsnative/volume space containing all specified areas (benson, wang atlas) and output corresponding information, apply this mask and convert the preprocessed surface/volume bold files to 2D NIFIT2 files as preparation for prfanalyze
-
link the correct stimulus appertures to the respective bold files by creating events.tsv
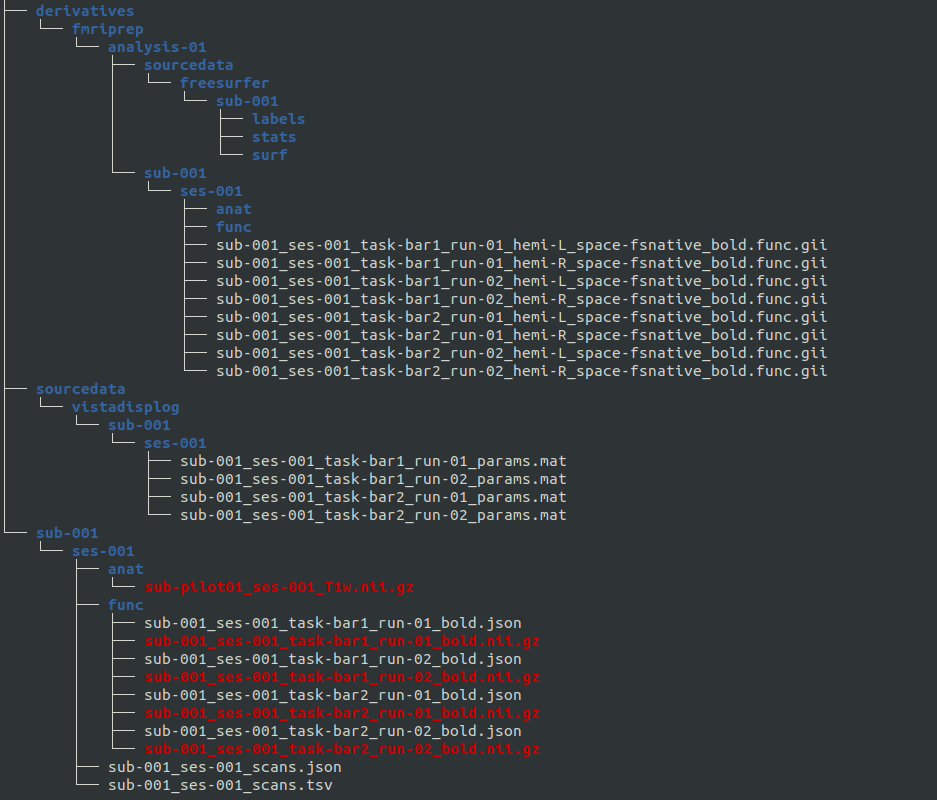
For running prfprepare you need to provide the following folder structure:
<BASE_DIR>/BIDS has to contain: -) the preprocessed data in fsnative space (for analysis_space=fsnative) or T1w space (for analysis_type=volume) and the corresponding freesurfer segmenataion (fMRIPrep output in BIDS/derivatives/fmriprep/analysis-XX; this can be set when calling the fMRIPrep)
-) the log file from vistadisp (in BIDS/derivatives/sourcedata/vistadisplog/sub-XX/ses-XX) and the stimulus images (in BIDS/derivatives/sourcedata/stimuli)
-) unprocessed subject data in BIDS-compatible format (e.g. from HeuDiConv) for the BIDS layout
We provide an example rundocker.sh script with all necessary bindings and an example_config.json file in this repository. example_config.json, * mandatory 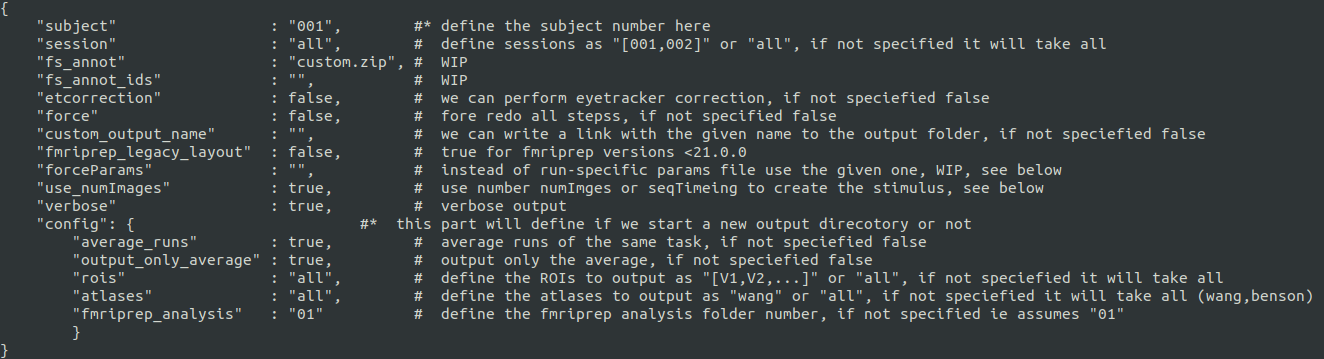
Subject and sessions can be defined as "all" (all that are found in BIDS folder), "001" for single or "[001,002,003]" for a list. Same works for rois. Set the analysisSpace either to "volume" for 3D analysis or "fsnative" for surface analysis.